

This project is a meeting place for users who share the R-Y93579 Y-DNA haplogroup, which means they are related along their paternal lines. Users in this group may want to share their family trees with each other to find overlaps and merge duplicate profiles in order to join or expand the World Family Tree and discover new relatives.
Y93579 is a deep subclade of the large R-U106 haplogroup.
R1b computed Jan 04 2023 using YFull v10.08.00
Total in System: 32620 samples, 57 ancient
R1b on YFull has 620 ancient or scientific study samples not in the FTDNA Big Y Tree
Y-DNA Next Generation Sequencing (NGS) is recommended over STR or individual SNP testing to confirm haplogroup R-U106 and downstream SNPs (NGS at least 15 Mbp at 30x depth). Whole Genome Sequencing (WGS) will also provide up to 100% Y-DNA coverage, but is not required.
atDNA testing can help locate additional candidates for testing.
It is recommended you share your BAM files with NGS Y-DNA scientists & researchers.
If you have an Autosomal DNA kit (atDNA), it is recommended you share/link your kit with relevant sites to locate additional matches.
Hamburg culture (15,500-13,100 ybp)

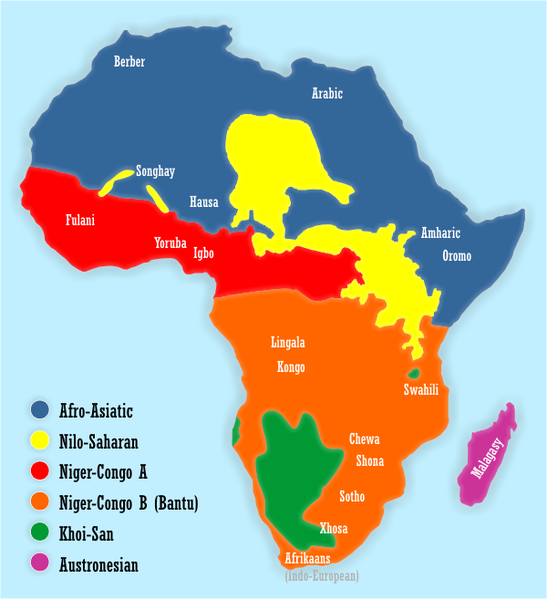

North African & Horn of Africa

Proto-Chadic Return to Africa



Wikipedia Haplogroup L-M20 (Elite Hun grave)

Indigenous Austronesians
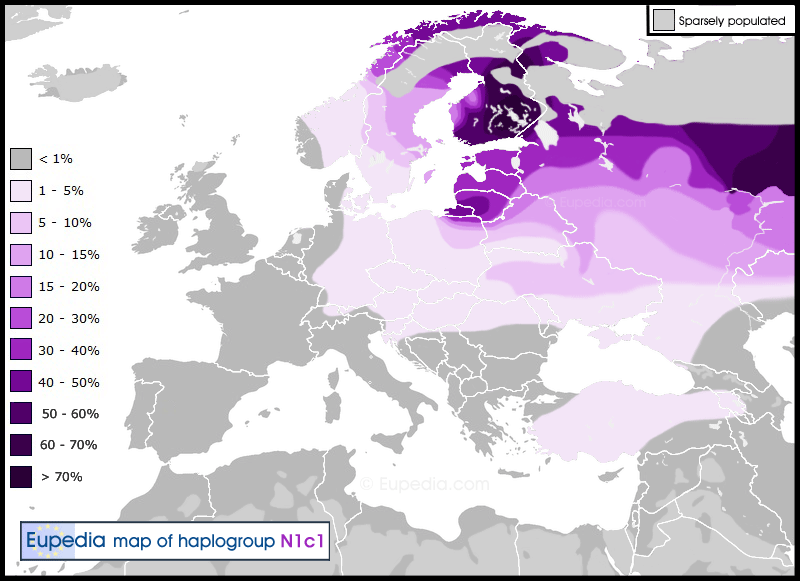

East Asian, Southeast Asian & Austronesian

Indigenous peoples of the Americas


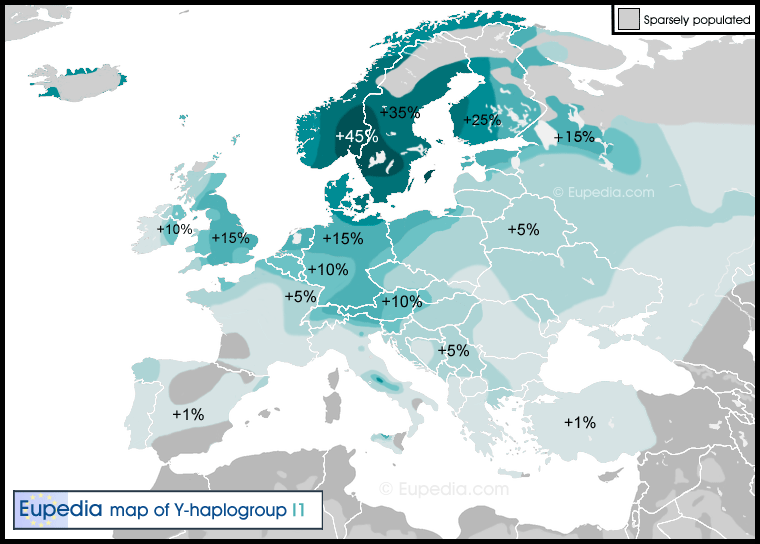
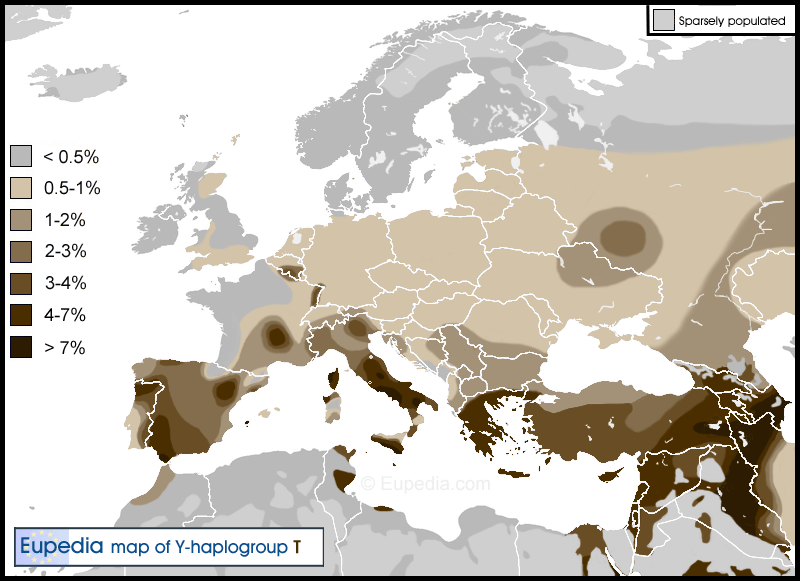
Ahrensburg culture (12,900-11,700 ybp)

Linear Pottery culture (circa 5500–4500 BCE)

Funnelbeaker culture (circa 4300-2800 BCE)
Swiderian culture (circa 11,000–8200 BCE)

Greeks

Proto-Indo-Europeans (R1)
Eupedia Yamna Culture (circa 3500-2500 BCE)

Several genetic studies performed since 2015 have given support to the Kurgan theory of Marija Gimbutas regarding the Indo-European Urheimat – that Indo-European languages spread throughout Europe from the Eurasian steppes and that the Yamnaya culture were Proto-Indo-Europeans. According to those studies, haplogroups R1b and R1a, now the most common in Europe (with R1a also being common in South Asia), would have expanded from the Pontic–Caspian steppes, along with the Indo-European languages. They also detected an autosomal component present in modern Europeans which was not present in Neolithic Europeans, which would have been introduced with paternal lineages R1b and R1a, as well as Indo-European languages in the Bronze Age.
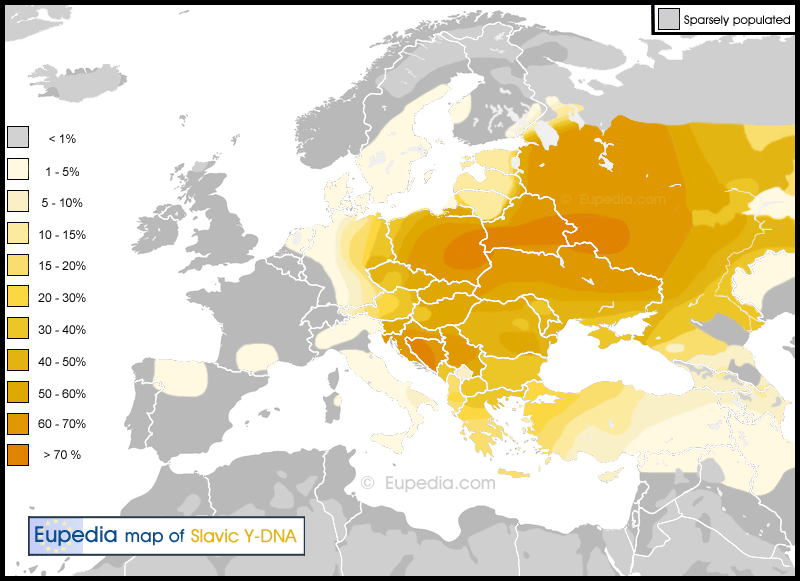
Proto-Baltio-Slavic (R1a-Z283)
Corded Ware culture (circa 2900-2350 BCE)
A genetic study conducted by Haak et al. (2015) found that a large proportion of the ancestry of the Corded Ware culture's population is similar to the Yamna culture, tracing the Corded Ware culture's origins to migrations of the Yamna from the steppes 4,500 years ago. About 75% of the DNA of late Neolithic Corded Ware skeletons found in Germany was a precise match to DNA from individuals of the Yamna culture. The same study estimated a 40–54% ancestral contribution of the Yamna in the DNA of modern Central & Northern Europeans, and a 20–32% contribution in modern Southern Europeans, excluding Sardinians (7.1% or less), and to a lesser extent Sicilians (11.6% or less). Haak et al. also note that their results "suggest" that haplogroups R1b and R1a "spread into Europe from the East after 3,000 BCE.
In terms of phenotypes, Wilde et al. (2014) and Haak et al. (2015) found that the intrusive Yamna population, generally inferred to be the first speakers of an Indo-European language in the Corded Ware culture zone, were overwhelmingly dark-eyed (brown), dark-haired and had a skin colour that was moderately light, though somewhat darker than that of the average modern European. These studies also showed that light pigmentation traits had already existed in pre-Indo-European Neolithic Europeans (in both farmers and hunter-gatherers), so long-standing philological attempts to correlate them with the arrival of Indo-Europeans from the steppes were misguided.
Autosomal DNA tests also indicate that the Yamna migration from the steppes introduced a component of ancestry referred to as "Ancient North Eurasian" admixture into Europe. "Ancient North Eurasian" is the name given in genetic literature to a component that represents descent from the people of the Mal'ta-Buret' culture or a population closely related to them. The "Ancient North Eurasian" genetic component is visible in tests of the Yamna people as well as modern-day Europeans, but not of Western or Central Europeans predating the Corded Ware culture.
Turkic (R1a-Z93)

Proto-Balkano-Anatolian-Italo-Celtic-Germanic (R1b-M269)
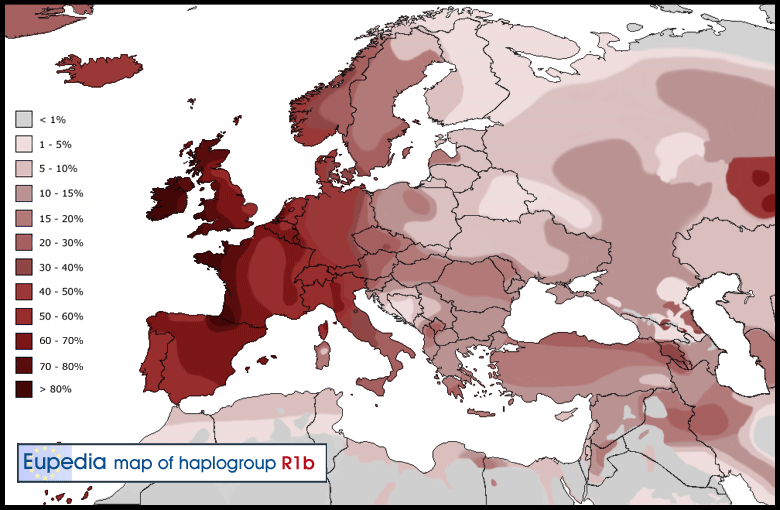
Balkano-Anatolian (R1b-Z2103)

Proto-Italo-Celtic-Germanic (R1b-L11)
Funnelbeaker culture ("circa" 4300-2800 BCE)
Genetic studies suggest that Funnelbeaker women were incorporated into the Corded Ware culture through intermixing with incoming Corded Ware males, and that people of the Corded Ware culture continued to use Funnelbeaker megaliths as burial grounds. Subsequent cultures of Late Neolithic, Bronze Age, and Iron Age Central Europe display strong maternal genetic affinity with the Funnelbeaker culture.
The evidence suggested that the Battle Axe culture entered Scandinavia through a migration from Eastern Europe, after which Battle Axe males mixed with Funnelbeaker females.
Early papers publishing results on European-wide Y-DNA marker frequencies, such as those of Semino (2000) and Rosser (2000), correlated haplogroup R1b-M269 with the earliest episodes of European colonization by anatomically modern humans (AMH). The peak frequencies of M269 in Iberia (especially the Basque region) and the Atlantic façade were postulated to represent signatures of re-colonization of the European West following the Last Glacial Maximum. However, even prior to recent criticisms and refinements, the idea that Iberian R1b carrying males repopulated most of western Europe was not consistent with findings which revealed that Italian M269 lineages are not derivative of Iberian ones.
More recently, data and calculations from Myres et al. (2011), Cruciani et al. (2011) Arredi et al. (2007), and Balaresque et al. (2010) suggest a Late Neolithic entry of M269 into Europe.
These hypotheses appear to be corroborated by more direct evidence from ancient DNA. R1b was detected in two male skeletons from a German Bell Beaker site dated to 2600–2500 BCE at Kromsdorf, one of which tested positive for M269 but negative for its U106 subclade (note that the P312 subclade was not tested for), while for the other skeleton the M269 test was unclear. A later Bell Beaker male skeleton from Quedlinburg, Germany dated to 2296–2206 BCE tested positive for R1b M269 P312 subclade. Ancient Y-DNA results for the remains of Beaker people from Iberia have yet to be obtained.

Tumulus culture (circa 1600 1200 BCE)
Proto-Italo-Celtic (R1b-P312)
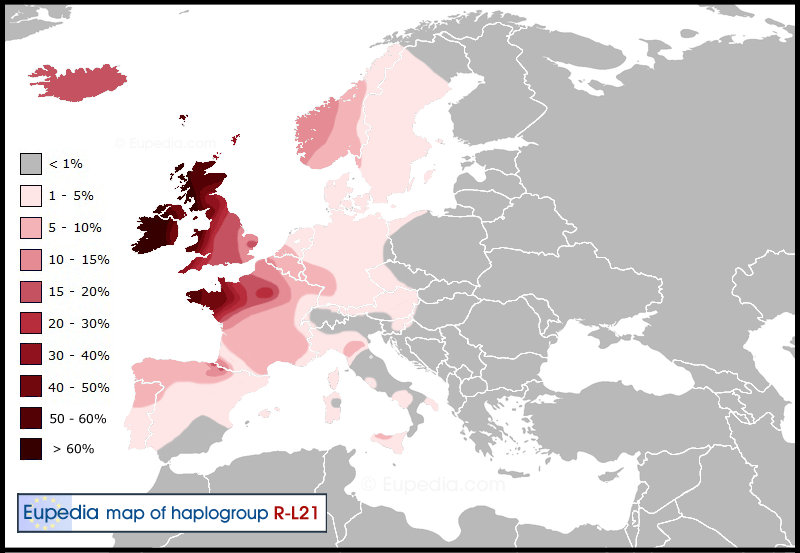
Atlantic Celts (R1b-L21)
Gallic & Iberian Celts (R1b-DF27)

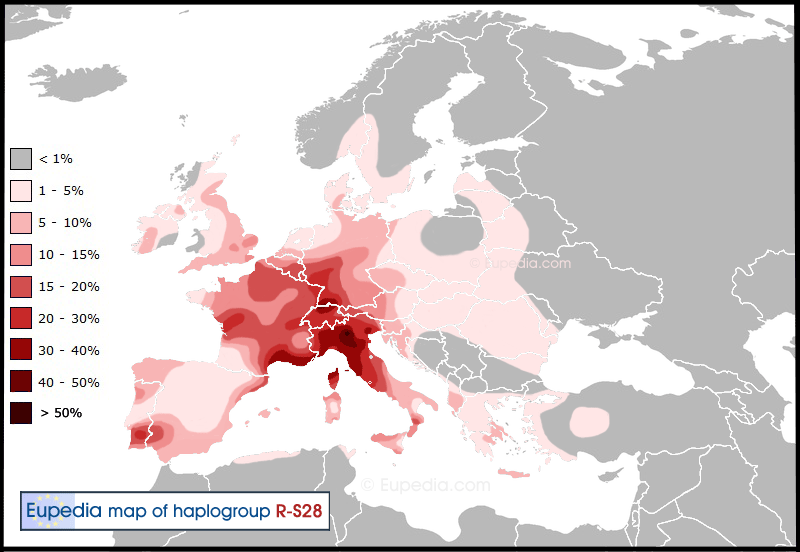
Alpine Celts (R1b-U152)
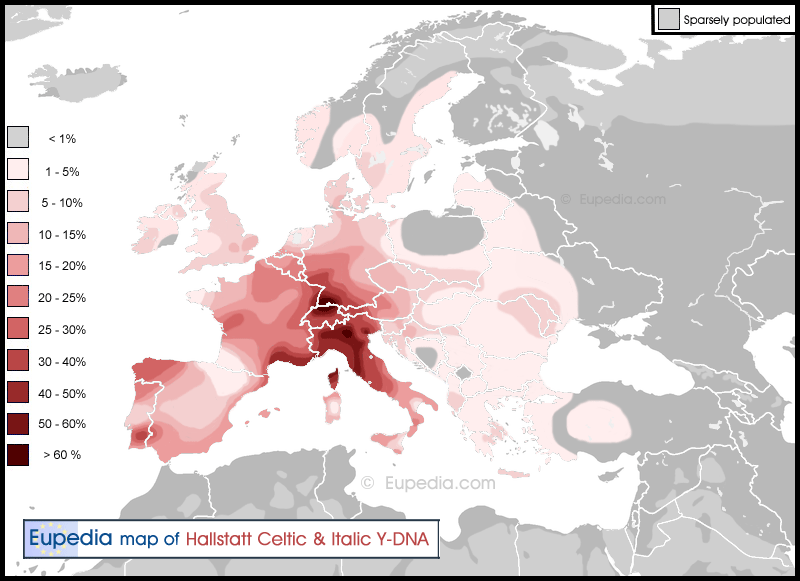
Terramare culture (circa 1700–1150 BCE)

Elp culture (circa 1800-800 BCE)